Authors
Lingsha Cheng, Haoqian Wu, Xiaoying Cai, Youying Zhang, Siqi Yu, Yuanlong Hou, Zhe Yin, Qingyuan Yan, Qiong Wang, Taipeng Sun, Guangji Wang, Yonggui Yuan, Xueli Zhang, Haiping Hao, Xiao Zheng
Lab
Journal
Cell Host & Microbe
Abstract
Summary
Gene-environment interactions shape behavior and susceptibility to depression. However, little is known about the signaling pathways integrating genetic and environmental inputs to impact neurobehavioral outcomes. We report that gut G-protein-coupled receptor, Gpr35, engages a microbe-to-brain metabolic pathway to modulate neuronal plasticity and depressive behavior in mice. Psychological stress decreases intestinal epithelial Gpr35, genetic deletion of which induces depressive-like behavior in a microbiome-dependent manner. Gpr35to/to mice and individuals with depression have increased Parabacteroides distasonis, and its colonization to wild-type mice induces depression. Gpr35to/to and Parabacteroides distasonis-colonized mice show reduced indole-3-carboxaldehyde (IAld) and increased indole-3-lactate (ILA), which are produced from opposing branches along the bacterial catabolic pathway of tryptophan. IAld and ILA counteractively modulate neuroplasticity in the nucleus accumbens, a brain region linked to depression. IAld supplementation produces anti-depressant effects in mice with stress or gut epithelial Gpr35 deficiency. Together, these findings elucidate a gut microbe-brain signaling mechanism that underlies susceptibility to depression.
Keywords/Topics
depressive ;disorder ;genetic risk ;Gpr35 ;gut microbiome; gut - brain axis ;tryptophan metabolism ;neural plasticity ;indole - 3 - carboxaldehyde
BIOSEB Instruments Used:
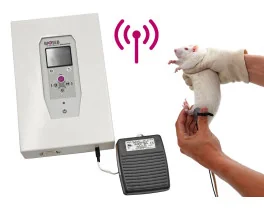
Electronic Von Frey - Wireless (BIO-EVF-WRS)
Source :
 Pain - Thermal Allodynia / Hyperalgesia
Pain - Thermal Allodynia / Hyperalgesia Pain - Spontaneous Pain - Postural Deficit
Pain - Spontaneous Pain - Postural Deficit Pain - Mechanical Allodynia / Hyperalgesia
Pain - Mechanical Allodynia / Hyperalgesia Learning/Memory - Attention - Addiction
Learning/Memory - Attention - Addiction Physiology & Respiratory Research
Physiology & Respiratory Research










![Dynamic Weight Bearing 2.0 – Postural Module [Add-on]](https://bioseb.com/733-home_default/dynamic-weight-bearing-20-add-on-postural-module.jpg)



















 Pain
Pain Central Nervous System (CNS)
Central Nervous System (CNS) Neurodegeneration
Neurodegeneration Sensory system
Sensory system Motor control
Motor control Mood Disorders
Mood Disorders Other disorders
Other disorders Muscular system
Muscular system Joints
Joints Metabolism
Metabolism Cross-disciplinary subjects
Cross-disciplinary subjects CONFERENCES & MEETINGS 2026
CONFERENCES & MEETINGS 2026 